
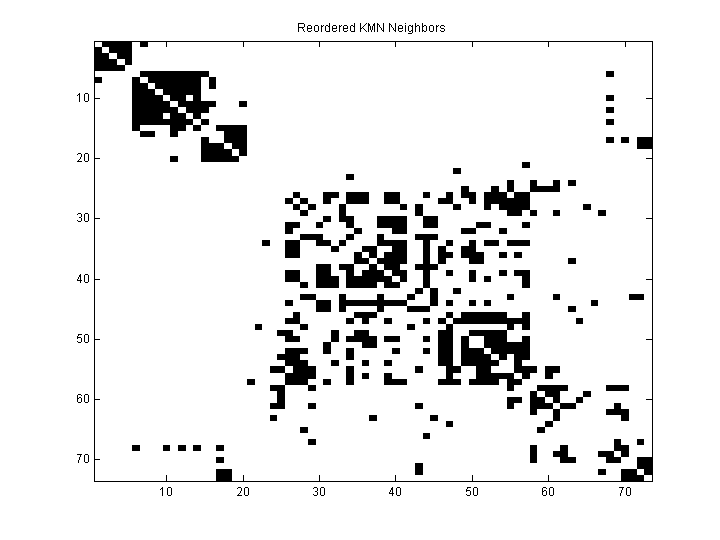
Each point is a row.





| 1 | 13 | S594 | 1437.cel | ||
| 2 | 16 | S593 S594 | 148.cel | ||
| 3 | 4 | S593 S594 | 1076.cel | ||
| 4 | 19 | S593 S594 | 1553.cel | ||
| 5 | 25 | S593 S594 | 1780.cel | ||
| 6 | 18 | S595 S597 S598 | 15.cel | ||
| 7 | 8 | S595 S597 S598 | 1308.cel | ||
| 8 | 11 | S595 S597 S598 | 1399.cel | ||
| 9 | 27 | S595 S597 S598 | 190.cel | ||
| 10 | 34 | S595 S597 S598 | 269.cel | ||
| 11 | 7 | S595 S597 S598 | 129.cel | ||
| 12 | 44 | S595 S597 S598 | 345.cel | ||
| 13 | 1 | S595 S597 S598 | 100R.cel | ||
| 14 | 61 | S595 S597 S598 | 57.cel | ||
| 15 | 37 | S597 S598 | 284.cel | ||
| 16 | 5 | S596 S598 | 12.cel | ||
| 17 | 40 | S596 S598 | 300.cel | ||
| 18 | 12 | S596 S598 | 1430.cel | ||
| 19 | 56 | S596 S598 | 528bis.cel | ||
| 20 | 62 | S596 S598 | 574.cel | ||
| 21 | 26 | S592 S596 S598 | 1854R.cel | ||
| 22 | 32 | S592 S596 S598 | 246.cel | ||
| 23 | 2 | S592 S596 S598 | 104.cel | ||
| 24 | 10 | S592 S596 S598 | 1360.cel | ||
| 25 | 14 | S592 S596 S598 | 1453.cel | ||
| 26 | 6 | S592 S596 S598 | 1284.cel | ||
| 27 | 15 | S592 S596 S598 | 147.cel | ||
| 28 | 21 | S592 S596 S598 | 157.cel | ||
| 29 | 42 | S592 S596 S598 | 328.cel | ||
| 30 | 23 | S592 S596 S598 | 1621.cel | ||
| 31 | 24 | S592 S596 S598 | 1622R.cel | ||
| 32 | 28 | S592 S596 S598 | 218.cel | ||
| 33 | 22 | S592 S596 S598 | 159.cel | ||
| 34 | 29 | S592 S596 S598 | 222.cel | ||
| 35 | 9 | S592 S596 S598 | 1317.cel | ||
| 36 | 30 | S592 S596 S598 | 236.cel | ||
| 37 | 50 | S592 S596 S598 | 428.cel | ||
| 38 | 39 | S592 S596 S598 | 30.cel | ||
| 39 | 49 | S592 S596 S598 | 396.cel | ||
| 40 | 46 | S592 S596 S598 | 356bis.cel | ||
| 41 | 71 | S592 S596 S598 | 97.cel | ||
| 42 | 31 | S592 S596 S598 | 242.cel | ||
| 43 | 45 | S592 S596 S598 | 349.cel | ||
| 44 | 51 | S592 S596 S598 | 444.cel | ||
| 45 | 52 | S592 S596 S598 | 458.cel | ||
| 46 | 20 | S592 S596 S598 | 155R.cel | ||
| 47 | 33 | S592 S596 S598 | 254.cel | ||
| 48 | 55 | S592 S596 S598 | 527.cel | ||
| 49 | 53 | S592 S596 S598 | 461.cel | ||
| 50 | 60 | S592 S596 S598 | 566.cel | ||
| 51 | 63 | S592 S596 S598 | 65R.cel | ||
| 52 | 35 | S592 S596 S598 | 274.cel | ||
| 53 | 41 | S592 S596 S598 | 323.cel | ||
| 54 | 67 | S592 S596 S598 | 84.cel | ||
| 55 | 17 | S592 S596 S598 | 1497R.cel | ||
| 56 | 68 | S592 S596 S598 | 90.cel | ||
| 57 | 70 | S592 S596 S598 | 94bis.cel | ||
| 58 | 38 | S592 S596 S598 | 288.cel | ||
| 59 | 54 | S592 S596 S598 | 506.cel | ||
| 60 | 72 | S592 S596 S598 | ge197.cel | ||
| 61 | 58 | S592 S596 S598 | 558.cel | ||
| 62 | 48 | S592 S596 S598 | 390.cel | ||
| 63 | 69 | S592 S596 S598 | 93.cel | ||
| 64 | 43 | S592 S596 S598 | 344.cel | ||
| 65 | 59 | S592 S596 S598 | 563.cel | ||
| 66 | 64 | S592 S596 S598 | 70.cel | ||
| 67 | 47 | S592 S596 S598 | 360.cel | ||
| 68 | 36 | S592 S596 S598 | 283.cel | ||
| 69 | 73 | S592 S596 S598 | ge205.cel | ||
| 70 | 65 | S596 S598 | 80.cel | ||
| 71 | 57 | S596 S598 | 547.cel | ||
| 72 | 3 | S596 S598 | 106.cel | ||
| 73 | 66 | S596 S598 | 83.cel |
| 1 | 22 | 204337_at | RGS4 Chr:1q23.2 | |
| 2 | 23 | 204339_s_at | RGS4 Chr:1q23.2 | |
| 3 | 16 | 203798_s_at | VSNL1 Chr:2p24.3 | |
| 4 | 93 | 228262_at | FLJ14503 Chr:xp22.13 | |
| 5 | 40 | 206376_at | SLC6A15 Chr:12q21.3 | |
| 6 | 91 | 227702_at | CYP4X1 Chr:1p33 | |
| 7 | 41 | 206456_at | GABRA5 Chr:15q11.2-q12 | |
| 8 | 139 | 243929_at | --- --- | |
| 9 | 72 | 219937_at | TRHDE Chr:12q15-q21 | |
| 10 | 63 | 214945_at | NY-REN-7 Chr:5q35.3 | |
| 11 | 8 | 1556366_s_at | FLJ33708 Chr:6p24.3 | |
| 12 | 69 | 219425_at | SULT4A1 Chr:22q13.2-q13.31 | |
| 13 | 3 | 1553864_at | DKFZp761H2121 Chr:10q26.13 | |
| 14 | 43 | 206678_at | GABRA1 Chr:5q34-q35 | |
| 15 | 113 | 233002_at | KIAA1622 Chr:14q32.2 | |
| 16 | 101 | 230137_at | FLJ30834 Chr:4q27 | |
| 17 | 73 | 220030_at | STYK1 Chr:12p13.31 | |
| 18 | 44 | 206780_at | GAD2 Chr:10p11.23 | |
| 19 | 7 | 1556351_at | HCN1 Chr:5p12 | |
| 20 | 5 | 1556095_at | UNC13C Chr:15q21.1 | |
| 21 | 19 | 204081_at | NRGN Chr:11q24 | |
| 22 | 56 | 213921_at | SST Chr:3q28 | |
| 23 | 32 | 205113_at | NEF3 Chr:8p21 | |
| 24 | 21 | 204229_at | SLC17A7 Chr:19q13 | |
| 25 | 135 | 242404_at | --- --- | |
| 26 | 1 | 1552714_at | CREG2 Chr:2q12.1 | |
| 27 | 108 | 231489_x_at | --- --- | |
| 28 | 37 | 205827_at | CCK Chr:3p22-p21.3 | |
| 29 | 140 | 244118_at | GABRA1 Chr:5q34-q35 | |
| 30 | 2 | 1553422_s_at | A2BP1 Chr:16p13.3 | |
| 31 | 9 | 1557122_s_at | --- --- | |
| 32 | 81 | 224650_at | MAL2 Chr:8q23 | |
| 33 | 18 | 203999_at | SYT1 Chr:12cen-q21 | |
| 34 | 50 | 210247_at | SYN2 Chr:3p25 | |
| 35 | 112 | 232426_at | --- --- | |
| 36 | 141 | 52837_at | KIAA1644 Chr:22q13 | |
| 37 | 74 | 220359_s_at | ARPP-21 Chr:3p22.1 | |
| 38 | 75 | 221217_s_at | A2BP1 Chr:16p13.3 | |
| 39 | 17 | 203998_s_at | SYT1 Chr:12cen-q21 | |
| 40 | 49 | 210227_at | DLGAP2 Chr:8p23 | |
| 41 | 6 | 1556096_s_at | UNC13C Chr:15q21.1 | |
| 42 | 105 | 230551_at | --- --- | |
| 43 | 120 | 237384_x_at | --- --- | |
| 44 | 119 | 236456_at | PTPN5 Chr:11p15.1 | |
| 45 | 132 | 241807_x_at | --- --- | |
| 46 | 35 | 205635_at | HAPIP Chr:3q21.2 | |
| 47 | 57 | 214044_at | RYR2 Chr:1q42.1-q43 | |
| 48 | 11 | 1568612_at | GABRG2 Chr:5q31.1-q33.1 | |
| 49 | 95 | 229039_at | SYN2 Chr:3p25 | |
| 50 | 25 | 204546_at | KIAA0513 Chr:16q24.1 | |
| 51 | 14 | 202507_s_at | SNAP25 Chr:20p12-p11.2 | |
| 52 | 71 | 219671_at | HPCAL4 Chr:1p34.2 | |
| 53 | 84 | 226086_at | SYT13 Chr:11p12-p11 | |
| 54 | 34 | 205414_s_at | KIAA0672 Chr:17p12 | |
| 55 | 70 | 219659_at | ATP8A2 Chr:13q12-13 | |
| 56 | 48 | 210040_at | SLC12A5 Chr:20q13.12 | |
| 57 | 130 | 241399_at | TAFA2 Chr:12q13.3-q14.1 | |
| 58 | 83 | 225111_s_at | NAPB Chr:20p12.3-p11.21 | |
| 59 | 127 | 239765_at | --- --- | |
| 60 | 86 | 227053_at | PACSIN1 Chr:6p21.3 | |
| 61 | 64 | 214956_at | --- --- | |
| 62 | 123 | 239357_at | --- --- | |
| 63 | 58 | 214098_at | KIAA1107 Chr:1p22.1 | |
| 64 | 62 | 214811_at | KIAA0318 Chr:12q24.33 | |
| 65 | 80 | 224209_s_at | GDA Chr:9q21.11-21.33 | |
| 66 | 36 | 205691_at | SYNGR3 Chr:16p13 | |
| 67 | 46 | 209726_at | CA11 Chr:19q13.3 | |
| 68 | 124 | 239579_at | ABHD7 Chr:1p22.1 | |
| 69 | 79 | 224164_at | TPM3 Chr:1q21.2 | |
| 70 | 96 | 229254_at | DKFZp761N1114 Chr:1q32.1 | |
| 71 | 131 | 241672_at | LOC400120 Chr:13q13.2 | |
| 72 | 65 | 215241_at | TMEM16C Chr:11p14.3 | |
| 73 | 13 | 202260_s_at | STXBP1 Chr:9q34.1 | |
| 74 | 39 | 206196_s_at | RPIP8 Chr:17q21.31 | |
| 75 | 77 | 223529_at | SYT4 Chr:18q12.3 | |
| 76 | 27 | 204722_at | SCN3B Chr:11q24.1 | |
| 77 | 60 | 214230_at | CDC42 Chr:1p36.1 | |
| 78 | 115 | 235066_at | MAP4 Chr:3p21 | |
| 79 | 78 | 223654_s_at | BRUNOL4 Chr:18q12 | |
| 80 | 122 | 238966_at | BRUNOL4 Chr:18q12 | |
| 81 | 47 | 210016_at | MYT1L Chr:2p25.3 | |
| 82 | 98 | 229300_at | --- --- | |
| 83 | 138 | 243666_at | BRUNOL4 Chr:18q12 | |
| 84 | 15 | 202712_s_at | CKMT1 Chr:15q15 | |
| 85 | 28 | 204723_at | SCN3B Chr:11q24.1 | |
| 86 | 103 | 230262_at | --- --- | |
| 87 | 136 | 242999_at | ARHGEF7 Chr:13q34 | |
| 88 | 45 | 209556_at | NCDN Chr:1p34.3 | |
| 89 | 85 | 226382_at | LOC283070 Chr:10p14 | |
| 90 | 10 | 1559545_at | --- --- | |
| 91 | 67 | 218623_at | HMP19 Chr:5q35.2 | |
| 92 | 100 | 230076_at | FLJ10156 Chr:17p13.2 | |
| 93 | 121 | 238786_at | --- --- | |
| 94 | 55 | 213904_at | --- --- | |
| 95 | 68 | 219308_s_at | AK5 Chr:1p31 | |
| 96 | 106 | 230698_at | --- --- | |
| 97 | 128 | 239767_at | --- --- | |
| 98 | 20 | 204134_at | PDE2A Chr:11q13.3 | |
| 99 | 4 | 1555958_at | CRTAC1 Chr:10q22 | |
| 100 | 24 | 204465_s_at | INA Chr:10q24.33 | |
| 101 | 38 | 206013_s_at | ACTL6 Chr:7q22 | |
| 102 | 30 | 204869_at | PCSK2 Chr:20p11.2 | |
| 103 | 61 | 214376_at | --- --- | |
| 104 | 110 | 232305_at | HMGCLL1 Chr:6p12.1 | |
| 105 | 118 | 236448_at | UNC5A Chr:5q35.3 | |
| 106 | 102 | 230255_at | GABRD Chr:1p36.3 | |
| 107 | 129 | 241382_at | --- --- | |
| 108 | 59 | 214111_at | OPCML Chr:11q25 | |
| 109 | 26 | 204697_s_at | CHGA Chr:14q32 | |
| 110 | 76 | 223500_at | CPLX1 Chr:4p16.3 | |
| 111 | 87 | 227226_at | C6orf117 Chr:6q15 | |
| 112 | 97 | 229294_at | JPH3 Chr:16q24.3 | |
| 113 | 107 | 230932_at | --- --- | |
| 114 | 104 | 230497_at | BRUNOL5 Chr:19p13 | |
| 115 | 133 | 242193_at | --- Chr:8q12.3 | |
| 116 | 125 | 239671_at | --- --- | |
| 117 | 31 | 204953_at | SNAP91 Chr:6q15 | |
| 118 | 88 | 227240_at | NGEF Chr:2q37 | |
| 119 | 90 | 227690_at | GABRB3 Chr:15q11.2-q12 | |
| 120 | 99 | 229724_at | GABRB3 Chr:15q11.2-q12 | |
| 121 | 111 | 232411_at | --- --- | |
| 122 | 116 | 235230_at | --- --- | |
| 123 | 114 | 233139_at | --- --- | |
| 124 | 117 | 236166_at | LOC285147 Chr:2p25.2 | |
| 125 | 12 | 202073_at | OPTN Chr:10p14 | |
| 126 | 33 | 205257_s_at | AMPH Chr:7p14-p13 | |
| 127 | 54 | 213307_at | SHANK2 Chr:11q13.2 | |
| 128 | 29 | 204763_s_at | GNAO1 Chr:16q13 | |
| 129 | 89 | 227641_at | MGC33974 Chr:16p13.3 | |
| 130 | 51 | 210721_s_at | PAK7 Chr:20p12 | |
| 131 | 92 | 227830_at | GABRB3 Chr:15q11.2-q12 | |
| 132 | 109 | 231986_at | RIMS1 Chr:6q12-q13 | |
| 133 | 137 | 243163_at | --- --- | |
| 134 | 52 | 212473_s_at | MICAL2 Chr:11p15.3 | |
| 135 | 42 | 206552_s_at | TAC1 Chr:7q21-q22 | |
| 136 | 94 | 229029_at | --- --- | |
| 137 | 53 | 213280_at | GARNL4 Chr:17p13.3 | |
| 138 | 66 | 216086_at | SV2C Chr:5q13.3 | |
| 139 | 82 | 224801_at | NDFIP2 Chr:13q22.2 | |
| 140 | 126 | 239678_at | --- --- | |
| 141 | 134 | 242344_at | --- --- |
| K (nearest neighbors) | 17 |
| Min T | 0.000000 |
| Max T | 0.250000 |
| Delta T | 0.004000 |
| Cycles | 3000 |
| Growth | TRUE |
| Stable delta T | 4 |
| Ignore dropout size | 1 |